Petteri Vainikka
Greetings from Athens!
My name is Petteri. I am from Finland, where I am currently finishing my MSc in chemistry.
I started my studies in the university of Turku in 2013, studying geology. After taking my introductory course in physical chemistry, I decided to pursue for a degree in chemistry (although I did finish my BSc in geology on the side because rocks are nice).
My initial plan of specializing in physical chemistry changed in 2015, when I joined David Palmer’s research group in Strathclyde University for a few months. After three exhilarating months of molecular dynamics (MD) and supercomputers, I knew there was no turning back. I returned in 2016, this time to learn the (very) basics of 3D-RISM and to deepen my knowledge of MD.
Since that, I’ve had the privilege of working in the MD-group in the university of Groningen (2017), and in the computational inorganic chemistry group in the university of Oxford (2018).
Late last January I received an email from PRACE, which advertised the Summer of HPC program. Although 91% of all the project topics looked alien to me, I could spot two topics related to computational chemistry. I decided to apply for project #14, named High throughput Virtual Screening to discover novel drug candidates. The motive here was quite simple: I had done a bit of classical stuff and a bit of non-classical stuff. Virtual screening was (and still is) an area where I have no experience whatsoever.
Late last March I got the good news from the program coordinator – I was selected. Fast-forward a few months, and I found myself in Edinburgh participating in the programs starting week, which was enjoyable. Made a few friends, learned quite a few new things about message-passing programming, and enjoyed a few nice local lagers in good company.
After the training week I flew to Athens, where I started working in Zoe Cournia’s group, based in the biomedical research foundation of the academy of Athens (BRFAA). During my stay here I will be mostly focusing on using virtual screening methods to try and find a isoform selective inhibitor for an enzyme known as ALDH7A1 (aldehyde dehydrogenase, seventh subfamily, member A1).
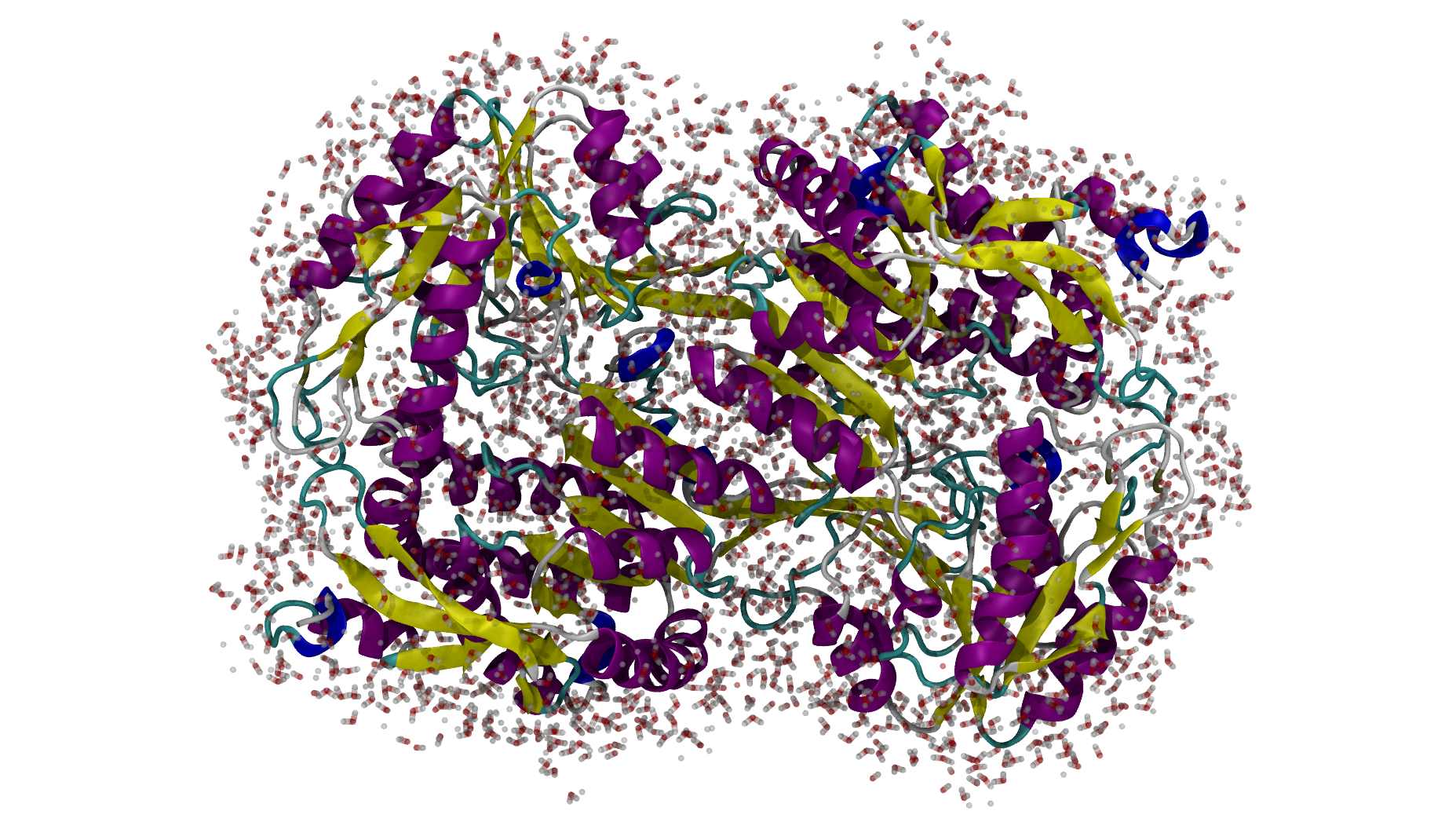
ALDH7A1 and its first solvation shell.
The motive for this project stems from the fact that overexpression of ALDH7A1 (with other isoforms such as ALDH1A1, ALDH1A3, ALDH2, and ALDH4A1) has been found in multiple types of cancer. In these cancers ALDHs promote drug resistance and amplify the amount of cancer stem cells (a group of cells which can initiate and propagate tumors, among other things).
To remain impartial, I should probably add that the ALDH family members also do non-cancerous things which include, but are not limited to, catalyzing reactions which detoxify reactive aldehydes in the body (such as acetaldehydes), and help reduce oxidative stress.
Stay tuned for more updates and cool figures!
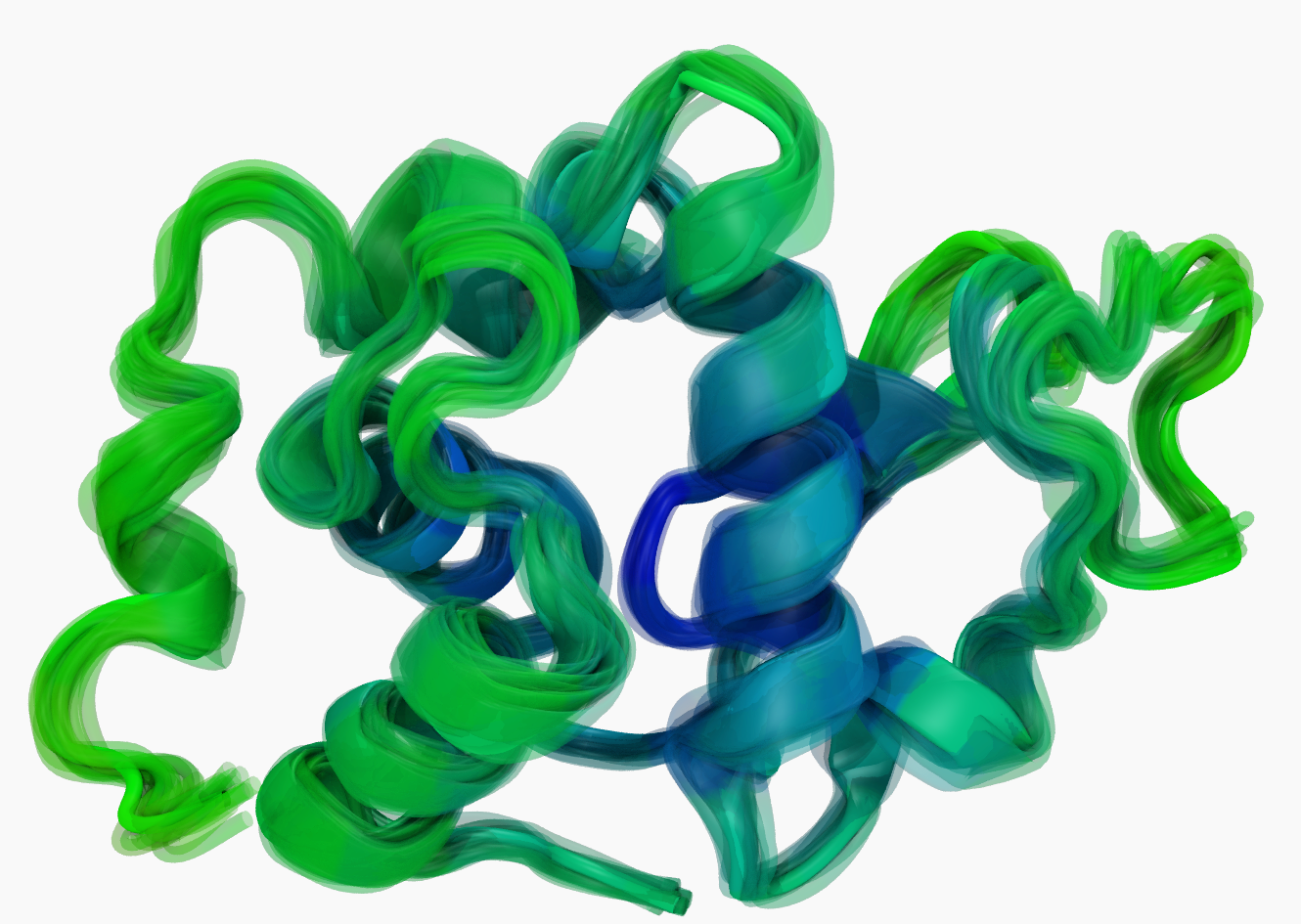

Hen egg-white lysozyme – just something from one of the tutorials I was working on earlier. The figure shows how the protein is moving throughout the simulation.
Further reading
- C. Chan, J. W.Y. Wong, C. Wong, M. K.L. Chan, and W. Fong: Human antiquitin: Structural and functional studies. Chem.-Biol. Interact. 2011, vol. 191 (1–3), p. 165-170.
- M. Luo, K. S. Gates, M. T. Henzl, and J. J. Tanner: Diethylaminobenzaldehyde is a covalent, irreversible inactivator of ALDH7A1. ACS Chem. Biol. 2015, vol. 10(3), p. 693-697.
- V. Vasiliou, D. C. Thompson, C. Smith, M. Fujita, and Y. Chen: Aldehyde dehydrogenases: From eye crystallins to metabolic disease and cancer stem cells. Chem.-Biol. Interact. 2013, vol. 202 (1–3), p. 2-10.

Leave a Reply