In Situ Visualization Technique
Traditionally, the process of performing numerical simulations consists of three separate steps: First, the input parameters (such as initial or boundary conditions) are specified, then the simulation is executed and finally, in order to determine the result, it is necessary to explore and analyse generated data. For several decades, increasing power of computer clusters and advancement in parallel computing methods allow scientists to perform more and more accurate simulations in various fields of human research.
On the other hand, increasing demands of the simulations need more data to be stored on a disk and analysed. However, the capabilities of computing environment which is responsible for transferring the data and communication have not grown up as rapid as the computational power. Dumping and processing of all the data calculated during the simulation would take too much time, so in practice this usually means that they are stored only at several time-steps or at much coarser resolution than the original data. The rest is just discarded and the significant part of information may be potentially lost.
In situ visualization stands for the technique, where the data are visualized in real-time, as it is being produced by simulation, and without involving storage resources. By using the visualization and simulation together, one can overcome the bottleneck of data transfer. Furthermore, this approach allows to monitor and interact with running simulation, so its parameters can be modified and scientist can immediately see the effect on investigating phenomena.
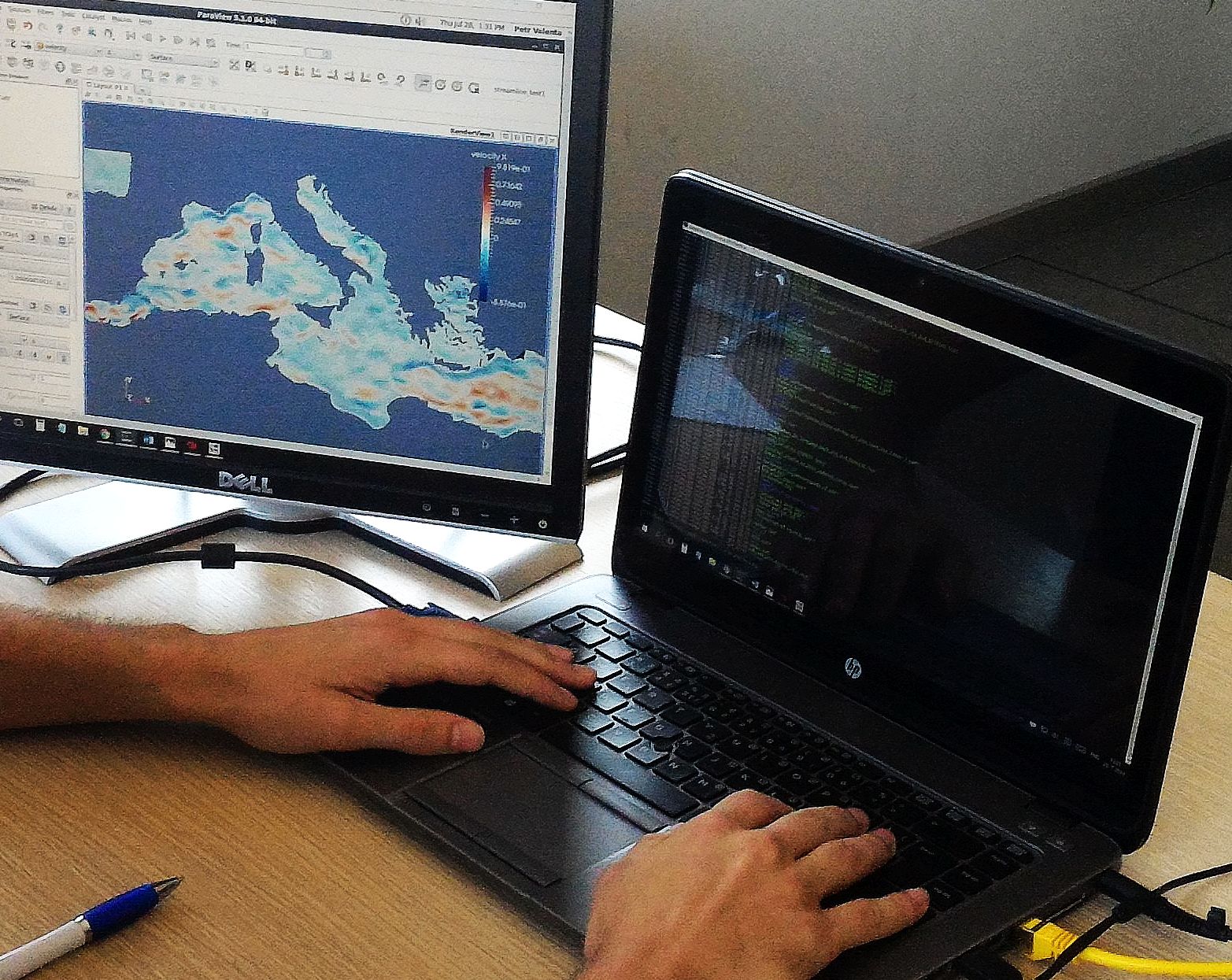

In situ visualization allows to observe results during the simulation runtime.
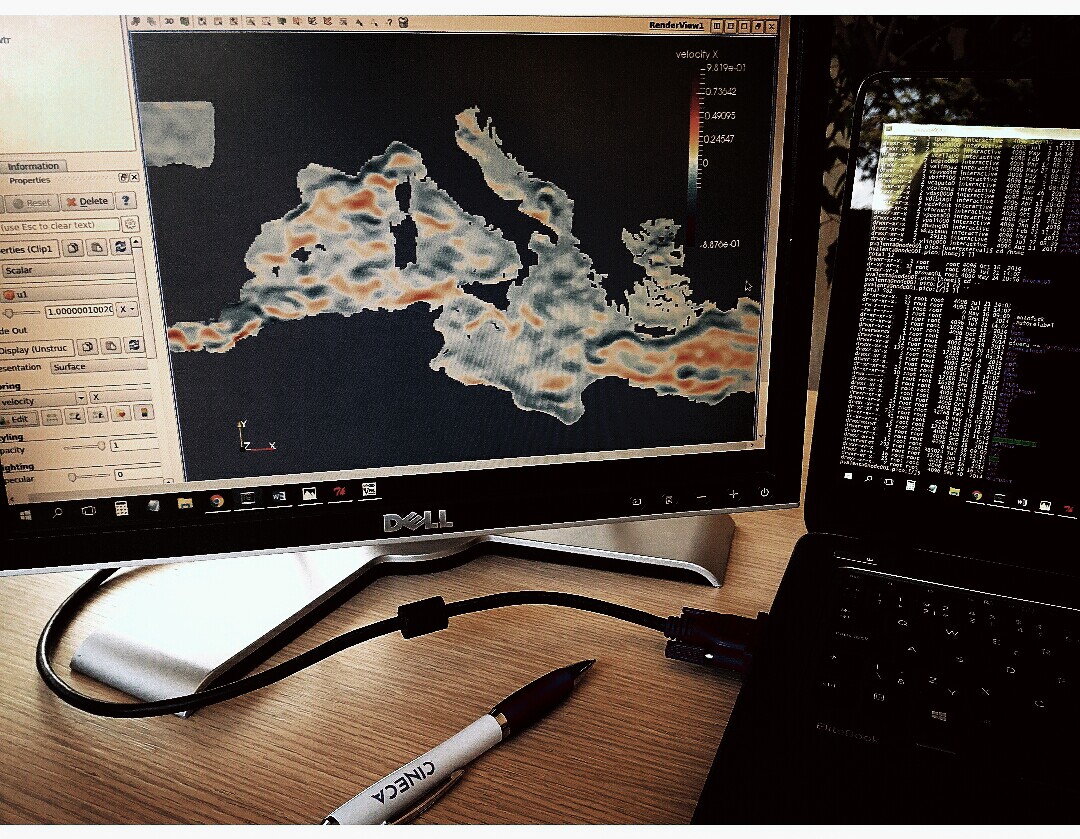
Researchers in OGS (National Institute of Oceanography and Experimental Geophysics) use a 3D numerical model OGSTM–BFM to study the nutrient and carbon cycles in the Mediterranean Sea and their sensitivity to climatic changes. The model computes biogeochemical fluxes which transform organic and inorganic components. Recent requirement of increasing the spatial and temporal scales of the simulations encounters aforementioned problems, thus the computation time is longer and it becomes very difficult to analyse produced data.
During the two months stay at CINECA supercomputing centre, I will be involved in a development of the 3D in situ visualization tool, which can be used to check and analyse the OGSTM–BFM model behaviour by consistently evaluating how the biogeochemical processes are influenced by the nutrient and carbon cycles, specifically related to the three main boundary conditions: the Atlantic inflow at Gibraltar Strait, the terrestrial inputs at rivers and the atmospheric deposition. The tool will be also beneficial to control the correctness of the computations during the simulation runtime. The implementation should be designed in such a way that allows portability to other coupled modelling systems used at OGS for many different purposes.
In general, in situ visualization creates an opportunity to explore and analyse much more data than is possible with traditional techniques. Also, as high performance computing moves towards the exascale era, in situ approach is widely predicted to become more and more important as an efficient tool for speed-up of large scale simulations.
